SPT6 promotes epidermal differentiation and blockade of an intestinal-like phenotype through control of transcriptional elongation | Nature Communications
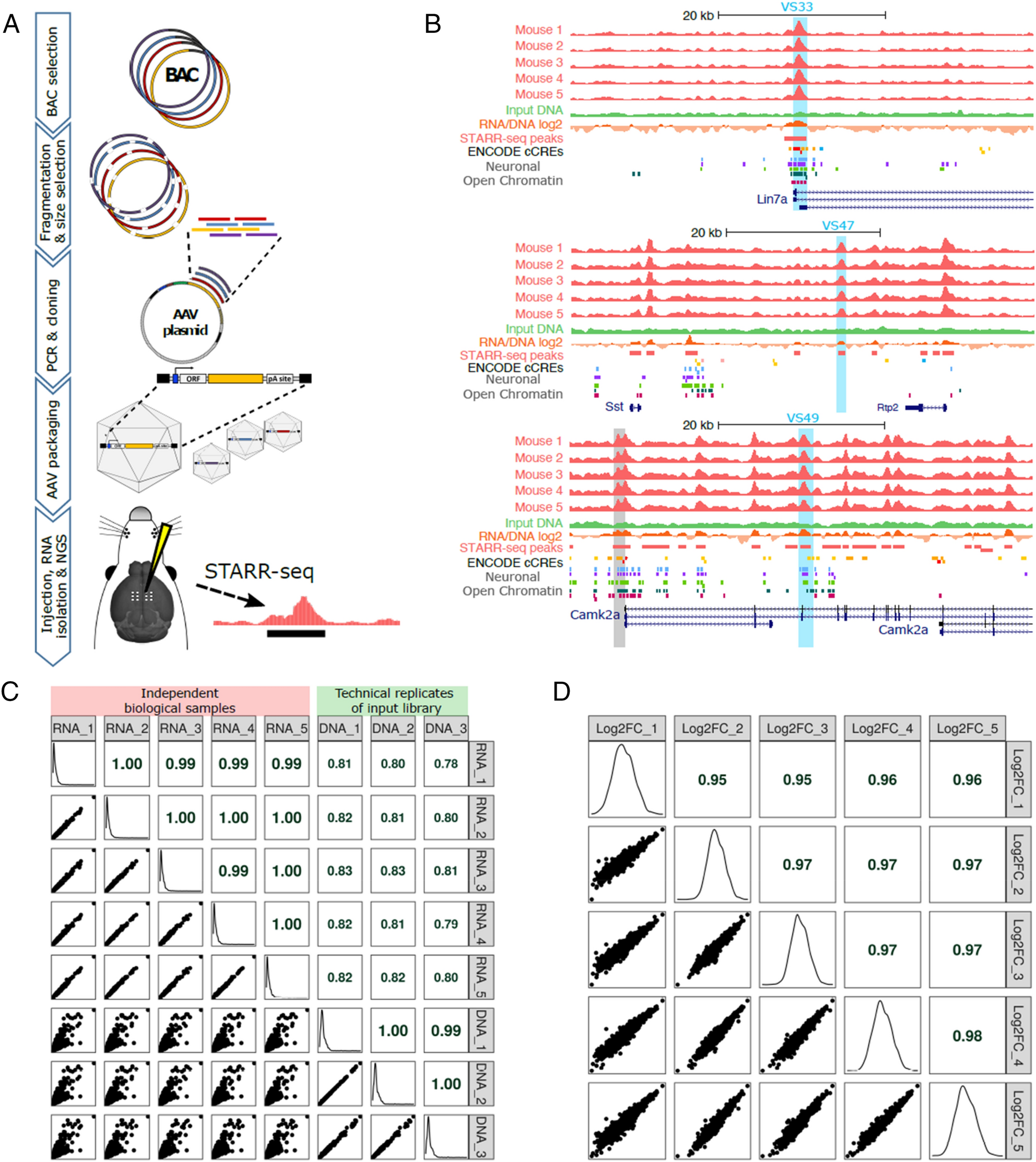
An unbiased AAV-STARR-seq screen revealing the enhancer activity map of genomic regions in the mouse brain in vivo | Scientific Reports

NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics

Discovery and validation of sub-threshold genome-wide association study loci using epigenomic signatures | eLife

CAGE-Seq Reveals that HIV-1 Latent Infection Does Not Trigger Unique Cellular Responses in a Jurkat T Cell Model | Journal of Virology

CAGE-Seq Reveals that HIV-1 Latent Infection Does Not Trigger Unique Cellular Responses in a Jurkat T Cell Model | Journal of Virology

Massively parallel characterization of transcriptional regulatory elements in three diverse human cell types | bioRxiv

CAGE-Seq Reveals that HIV-1 Latent Infection Does Not Trigger Unique Cellular Responses in a Jurkat T Cell Model | Journal of Virology

NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics

NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics

Massively parallel characterization of transcriptional regulatory elements in three diverse human cell types | bioRxiv

NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics

Massively parallel characterization of transcriptional regulatory elements in three diverse human cell types | bioRxiv

High-resolution annotation of the mouse preimplantation embryo transcriptome using long-read sequencing | Nature Communications

Orphan CpG islands boost the regulatory activity of poised enhancers and dictate the responsiveness of their target genes | bioRxiv

Orphan CpG islands boost the regulatory activity of poised enhancers and dictate the responsiveness of their target genes | bioRxiv